Project Associate Professor Kosuke Fujimoto of the Institute of Medical Science utilizes next-generation sequencers and supercomputers for genomic analysis of the gut microbiota.
An expert on gut viruses known as bacteriophages, he offers a primer on his field and research, which he conducts with world-class analytical tools and techniques at his disposal.
Genomic immunology and feces
Healthy human feces restore intestinal function in patients
FUJIMOTO Kosuke
Project Associate Professor
The Institute of Medical Science

More than 100 species of bacteria reside in our intestines, with their total number estimated to be around 100 trillion. This diverse gut microbiota is essential for human survival, producing metabolites essential for metabolism, and turning food into nutrients through digestion. At our laboratory, we are engaged in genomic analysis of these gut microorganisms.
Fecal transplantation guts infectious diseases
Advances in genomic analysis technologies, such as next-generation sequencers, have revealed that disruption and imbalance in the gut microbiota increase the risk of various diseases. This led to the concept of fecal transplantation, which involves administering healthy human gut microbiota to patients. A brown, filtered liquid derived from feces is introduced into the intestines using an endoscope or colonoscope. Although still at the clinical research stage in Japan, this procedure is performed as a standard treatment in Europe and the United States. Some countries have fecal banks, and there exist highly effective “super donor” feces.
Fecal microbiota treatment is particularly effective against Clostridioides difficile (C. difficile) infection. C. difficile bacteria are present in very small numbers in the intestines of healthy individuals. They usually remain dormant but produce toxins and cause enteritis, an inflammation of the intestines, when the gut microbiota is disrupted, for example by antibiotic use. In many cases, discontinuing use of the causative antibiotics leads to recovery; however, cases involving infections with highly virulent strains do not resolve easily. In the United States, 200,000 to 300,000 people contract this infection annually, with tens of thousands of deaths per year. Fecal transplantation works well against this condition, with some reports indicating that about 90% of patients have been successfully treated through this method. However, its precise mechanism remains unclear. Although feces from healthy donors are used, there is no established definition of what constitutes “normal feces.”
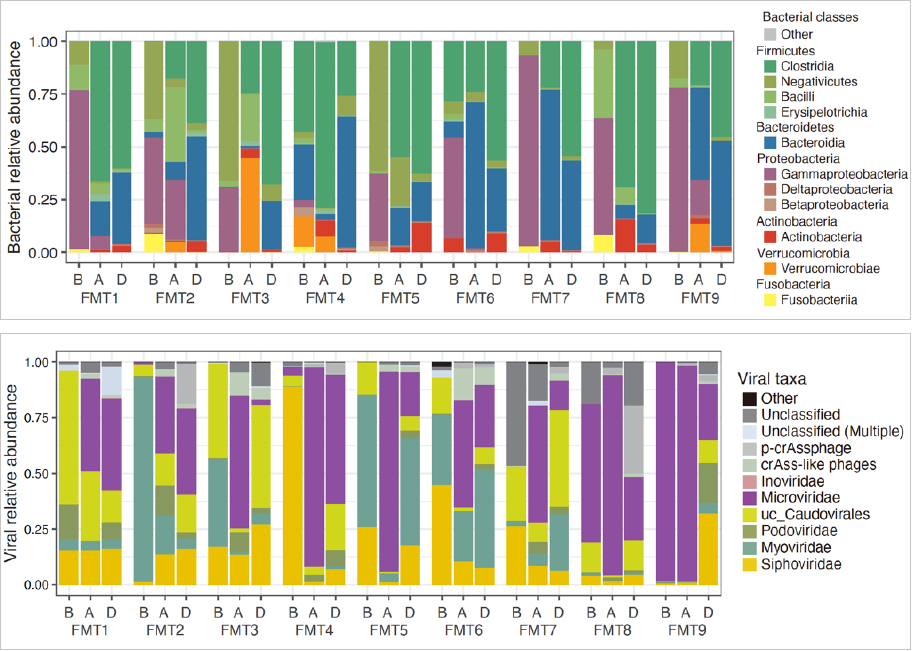
In our laboratory, we performed genomic analysis of gut bacteria and gut viruses using fecal samples from patients before and after fecal transplantation therapy, as well as those from donors. We found not only does the gut microbiota composition after transplantation become similar to that of the donor, but the functional genes carried by the gut microbiota turn alike as well. This clearly demonstrates the recovery of gut microbiota function.
Phage therapy makes comeback as hope against multidrug-resistant bacteria
Currently, we are focusing on genomic analysis of gut viruses called bacteriophages. Phages parasitize bacteria. Before the advent of antibiotics, there were efforts to use phages to kill target bacteria and treat infections. This phage therapy has regained attention due to the emergence of multidrug-resistant bacteria that are unresponsive to antibiotics.
Over the past decade, we have employed various methods to conduct genomic analysis of these phages. One of our achievements is extracting gut bacteria and phages from fecal samples of 101 healthy Japanese individuals, performing whole-genome analysis and constructing a database. This enables us to identify, based solely on genomic data, which phages can infect which bacteria without the need for bacterial culturing. Using the expertise we have accumulated so far, we aim to create innovative phage therapies and translate them into treatments for various diseases.